Simulate Covariance Matrices from Prior
rprior.RdSimulates covariance matrices from prior as specified in a MCMCglmm prior
Details
If alpha.V is NULL the distribution is an inverse-Wishart distribution (i.e. inverse-gamma in the univariate case). If alpha.V is non-null, MCMCglmm uses parameter expansion and the distribution is an inverse-Wishart mixture with no closed form density function expect in the univariate case (scaled F with 1 numerator degree of freedom).
Author
Jarrod Hadfield j.hadfield@ed.ac.uk
Examples
prior<-resolve_prior(F(10, 20), k=3, vtype="us")
# 3x3 covariance matrix with scaled (20) central F_{1,10} marginal variances
V<-rprior(2000, prior)
hist(V[,1], freq=FALSE, breaks=100, main="", xlab="Variance")
x<-seq(1e-6, max(V[,1]), length=1000)
mprior<-resolve_prior(F(10, 20), k=1, vtype="us")
# univariate prior for central F_{1,10} with scale 20
lines(dprior(x, mprior)~x)
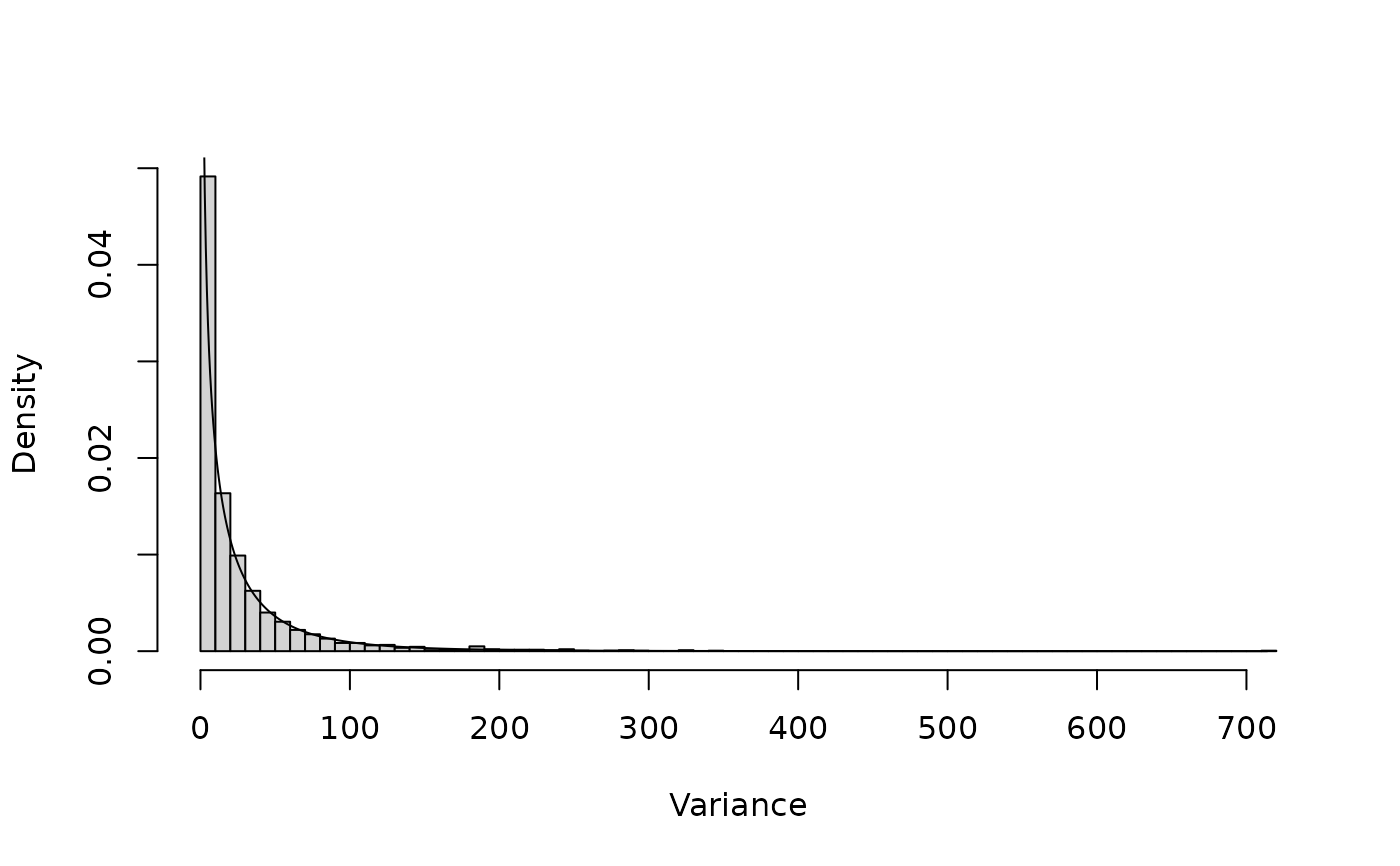 # Density for variance
# Density for variance